MICROBIAL MOLECULAR BIOLOGY & GENETICS
A) Genes: Structure, Replication & Mutation
Nucleic Acid Structure:
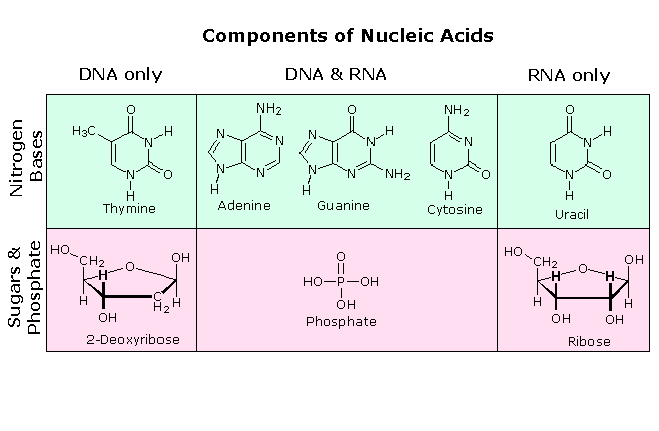
The structure and synthesis of purine and pyrimidine nucleotides as you all know. These nucleotides can be combined to form two types of nucleic acids; Deoxyribonucleic acid(DNA) & Ribonucleic acid( RNA). Ribonucleic acid is composed of the ribonucleosides of adenine, guanine, cytosine, and uracil (instead of thymine). In both DNA and
RNA, nucleosides are joined by phosphate groups to form long polynucleotides.
Nucleic acids are required for the storage and expression of genetic
information. There are two chemically distinct types of nucleic acids:
deoxyribonucleic acid (DNA) and ribonucleic acid . DNA, the repository of genetic information, is present not only in
chromosomes in the nucleus of eukaryotic organisms, but also in
mitochondria and the chloroplasts of plants. Prokaryotic cells, which
lack nuclei, have a single chromosome, but may also contain
nonchromosomal DNA in the form of plasmids. The genetic information
found in DNA is copied and transmitted to daughter cells through
DNA replication. The DNA contained in a fertilized egg encodes the
information that directs the development of an organism. This development
may involve the production of billions of cells. Each cell is
specialized, expressing only those functions that are required for it to
perform its role in maintaining the organism. Therefore, DNA must be
able to not only replicate precisely each time a cell divides, but also
to have the information that it contains be selectively expressed.
C) Linear & Circular DNA molecules:
Each chromosome in the nucleus of a eukaryote contains one long,
linear molecule of dsDNA, which is bound to a complex mixture of
proteins (histone and non-histone,) to form chromatin.
Eukaryotes have closed, circular DNA molecules in their mitochondria,
as do plant chloroplasts. A prokaryotic organism typically contains
a single, double-stranded, supercoiled, circular chromosome.
Each prokaryotic chromosome is associated with non-histone proteins
that can condense the DNA to form a nucleoid. In addition,
most species of bacteria also contain small, circular, extrachromosomal
DNA molecules called plasmids. Plasmid DNA carries genetic
information, and undergoes replication that may or may not be synchronized
to chromosomal division.
Note: Plasmids may carry genes that convey antibiotic
resistance to the host bacterium, and may
facilitate the transfer of genetic information from
one bacterium to another.
The Organization of DNA in Cells:
Although DNA exists as a double helix in both procaryotic and
eucaryotic cells, its organization differs in the two cell types.
DNA is organized in the form of a closed circle in almost
all procaryotes (the chromosome of Borrelia is a linear
DNA molecule). This circular double helix is further twisted into
supercoiled DNA and is associated with basic proteins
but not with the histones found complexed with almost all
eucaryotic DNA. These histone like proteins do appear to help organize
bacterial DNA into a coiled chromatin like structure.
DNA is much more highly organized in eucaryotic chromatin
and is associated with a variety of proteins, the
most prominent of which are histones. These are small, basic proteins
rich in the amino acids lysine and/or arginine. There are five
types of histones in almost all eucaryotic cells studied: H1, H2A,
H2B, H3, and H4. Eight histone molecules (two each of H2A,
H2B, H3, and H4) form an ellipsoid about 11 nm long and 6.5 to 7
nm in diameter . DNA coils around the surface of the
ellipsoid approximately 13
4 turns or 166 base pairs before proceeding
on to the next. This complex of histones plus DNA is called a nucleosome. Thus DNA gently isolated from chromatin looks like
a string of beads. The stretch of DNA between the beads or nucleosomes,
the linker region, varies in length from 14 to over 100 base
pairs. Histone H1 appears to associate with the linker regions to aid
the folding of DNA into more complex chromatin structures . When folding reaches a maximum, the chromatin takes
the shape of the visible chromosomes seen in eucaryotic cells during
mitosis and meiosis.
Cited By Kamal Singh Khadka
Msc Microbiology, TU
Assistant Professor in PU, RE-COST, PNC NA, LA
POKHARA, NEPAL
Nucleic Acid Structure:
The structure and synthesis of purine and pyrimidine nucleotides as you all know. These nucleotides can be combined to form two types of nucleic acids; Deoxyribonucleic acid(DNA) & Ribonucleic acid( RNA). Ribonucleic acid is composed of the ribonucleosides of adenine, guanine, cytosine, and uracil (instead of thymine). In both DNA and
RNA, nucleosides are joined by phosphate groups to form long polynucleotides.
Nucleic acids are required for the storage and expression of genetic
information. There are two chemically distinct types of nucleic acids:
deoxyribonucleic acid (DNA) and ribonucleic acid . DNA, the repository of genetic information, is present not only in
chromosomes in the nucleus of eukaryotic organisms, but also in
mitochondria and the chloroplasts of plants. Prokaryotic cells, which
lack nuclei, have a single chromosome, but may also contain
nonchromosomal DNA in the form of plasmids. The genetic information
found in DNA is copied and transmitted to daughter cells through
DNA replication. The DNA contained in a fertilized egg encodes the
information that directs the development of an organism. This development
may involve the production of billions of cells. Each cell is
specialized, expressing only those functions that are required for it to
perform its role in maintaining the organism. Therefore, DNA must be
able to not only replicate precisely each time a cell divides, but also
to have the information that it contains be selectively expressed.
STRUCTURE OF DNA:
DNA is a polymer of deoxyribonucleoside monophosphates covalently
linked by 3'→5'–phosphodiester bonds. With the exception of a few
viruses that contain single-stranded (ss) DNA, DNA exists as a doublestranded
(ds) molecule, in which the two strands wind around each
other, forming a double helix. In eukaryotic cells, DNA is found associated
with various types of proteins (known collectively as nucleoprotein)
present in the nucleus, whereas in prokaryotes, the protein–DNA complex
is present in a nonmembrane-bound region known as the nucleoid.
A) 3'→5'-Phosphodiester bonds
Phosphodiester bonds join the 3'-hydroxyl group of the deoxy pentose
of one nucleotide to the 5'-hydroxyl group of the deoxy pentose of an
adjacent nucleotide through a phosphate group . The
resulting long, unbranched chain has polarity, with both a 5'-end (the
end with the free phosphate) and a 3'-end (the end with the free
hydroxyl) that are not attached to other nucleotides. The bases
located along the resulting deoxy ribose–phosphate backbone are, by
convention, always written in sequence from the 5'-end of the chain to the 3'-end.
B. Double helix:
In the double helix, the two chains are coiled around a common axis
called the axis of symmetry. The chains are paired in an anti parallel
manner, that is, the 5'-end of one strand is paired with the 3'-end of the other strand.
In the DNA helix, the hydrophilic
deoxyribose–phosphate backbone of each chain is on the outside of
the molecule, whereas the hydrophobic bases are stacked inside.
The overall structure resembles a twisted ladder. The spatial relationship
between the two strands in the helix creates a major (wide)
groove and a minor (narrow) groove. These grooves provide access
for the binding of regulatory proteins to their specific recognition
sequences along the DNA chain. Certain anticancer drugs, such as
dactinomycin (actinomycin D), exert their cytotoxic effect by intercalating
into the narrow groove of the DNA double helix, thus interfering
with DNA and RNA synthesis.
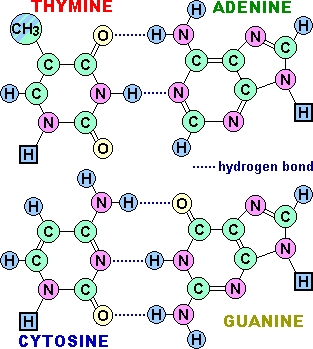
1) Base pairing:
The bases of one strand of DNA are paired with the
bases of the second strand, so that an adenine is always paired
with a thymine and a cytosine is always paired with a guanine.
[Note: The base pairs are perpendicular to the axis of the helix. Therefore, one polynucleotide chain of the DNA
double helix is always the complement of the other. Given the
sequence of bases on one chain, the sequence of bases on the
complementary chain can be determined [Note: The
specific base pairing in DNA leads to the Chargaff Rule: In any
sample of dsDNA, the amount of adenine equals the amount of
thymine, the amount of guanine equals the amount of cytosine, and
the total amount of purines equals the total amount of pyrimidines.]
The base pairs are held together by hydrogen bonds: two between
A and T and three between G and C. These hydrogen
bonds, plus the hydrophobic interactions between the stacked
bases, stabilize the structure of the double helix.
2) Separation of the two DNA strands in the double helix:
The two
strands of the double helix separate when hydrogen bonds
between the paired bases are disrupted. Disruption can occur in
the laboratory if the pH of the DNA solution is altered so that the
nucleotide bases ionize, or if the solution is heated. [Note:
Phosphodiester bonds are not broken by such treatment.] When
DNA is heated, the temperature at which one half of the helical
structure is lost is defined as the melting temperature (Tm). The
loss of helical structure in DNA, called denaturation, can be monitored
by measuring its absorbance at 260 nm. [Note: ssDNA has a
higher relative absorbance at this wavelength than does dsDNA.]
Because there are three hydrogen bonds between G and C but
only two between A and T, DNA that contains high concentrations
of A and T denatures at a lower temperature than G- and C-rich
DNA . Under appropriate conditions, complementary
DNA strands can reform the double helix by the process called
renaturation (or reannealing).
C) Linear & Circular DNA molecules:
Each chromosome in the nucleus of a eukaryote contains one long,
linear molecule of dsDNA, which is bound to a complex mixture of
proteins (histone and non-histone,) to form chromatin.
Eukaryotes have closed, circular DNA molecules in their mitochondria,
as do plant chloroplasts. A prokaryotic organism typically contains
a single, double-stranded, supercoiled, circular chromosome.
Each prokaryotic chromosome is associated with non-histone proteins
that can condense the DNA to form a nucleoid. In addition,
most species of bacteria also contain small, circular, extrachromosomal
DNA molecules called plasmids. Plasmid DNA carries genetic
information, and undergoes replication that may or may not be synchronized
to chromosomal division.
Note: Plasmids may carry genes that convey antibiotic
resistance to the host bacterium, and may
facilitate the transfer of genetic information from
one bacterium to another.
The Organization of DNA in Cells:
Although DNA exists as a double helix in both procaryotic and
eucaryotic cells, its organization differs in the two cell types.
DNA is organized in the form of a closed circle in almost
all procaryotes (the chromosome of Borrelia is a linear
DNA molecule). This circular double helix is further twisted into
supercoiled DNA and is associated with basic proteins
but not with the histones found complexed with almost all
eucaryotic DNA. These histone like proteins do appear to help organize
bacterial DNA into a coiled chromatin like structure.
DNA is much more highly organized in eucaryotic chromatin
and is associated with a variety of proteins, the
most prominent of which are histones. These are small, basic proteins
rich in the amino acids lysine and/or arginine. There are five
types of histones in almost all eucaryotic cells studied: H1, H2A,
H2B, H3, and H4. Eight histone molecules (two each of H2A,
H2B, H3, and H4) form an ellipsoid about 11 nm long and 6.5 to 7
nm in diameter . DNA coils around the surface of the
ellipsoid approximately 13
4 turns or 166 base pairs before proceeding
on to the next. This complex of histones plus DNA is called a nucleosome. Thus DNA gently isolated from chromatin looks like
a string of beads. The stretch of DNA between the beads or nucleosomes,
the linker region, varies in length from 14 to over 100 base
pairs. Histone H1 appears to associate with the linker regions to aid
the folding of DNA into more complex chromatin structures . When folding reaches a maximum, the chromatin takes
the shape of the visible chromosomes seen in eucaryotic cells during
mitosis and meiosis.
Cited By Kamal Singh Khadka
Msc Microbiology, TU
Assistant Professor in PU, RE-COST, PNC NA, LA
POKHARA, NEPAL




Comments